Cobalt »
PDB 6vyh-7biz »
6w54 »
Cobalt in PDB 6w54: Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans
Protein crystallography data
The structure of Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans, PDB code: 6w54
was solved by
M.Zeug,
N.Marckovic,
C.V.Iancu,
J.Tripp,
M.Oreb,
J.Choe,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 41.70 / 1.50 |
| Space group | H 3 |
| Cell size a, b, c (Å), α, β, γ (°) | 81.971, 81.971, 103.055, 90, 90, 120 |
| R / Rfree (%) | 14 / 16.6 |
Other elements in 6w54:
The structure of Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans also contains other interesting chemical elements:
| Potassium | (K) | 1 atom |
Cobalt Binding Sites:
The binding sites of Cobalt atom in the Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans
(pdb code 6w54). This binding sites where shown within
5.0 Angstroms radius around Cobalt atom.
In total only one binding site of Cobalt was determined in the Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans, PDB code: 6w54:
In total only one binding site of Cobalt was determined in the Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans, PDB code: 6w54:
Cobalt binding site 1 out of 1 in 6w54
Go back to
Cobalt binding site 1 out
of 1 in the Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans
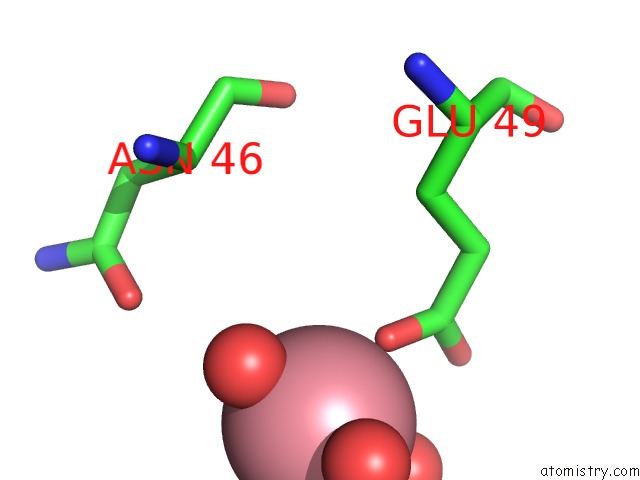
Mono view
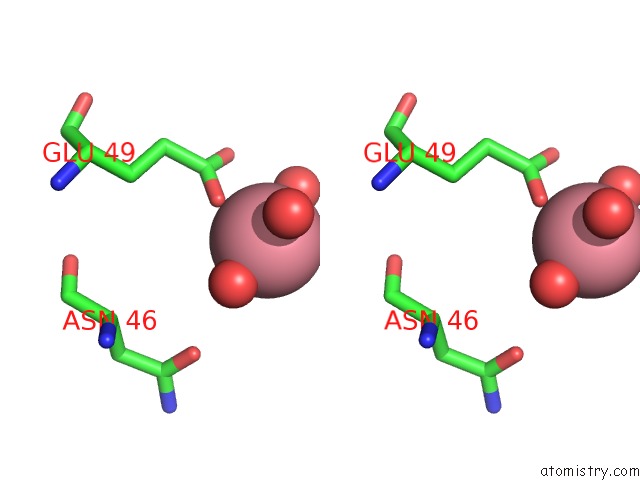
Stereo pair view

Mono view

Stereo pair view
A full contact list of Cobalt with other atoms in the Co binding
site number 1 of Crystal Structure of Gallic Acid Decarboxylase From Arxula Adeninivorans within 5.0Å range:
|
Reference:
M.Zeug,
N.Markovic,
C.V.Iancu,
J.Tripp,
M.Oreb,
J.Y.Choe.
Crystal Structures of Non-Oxidative Decarboxylases Reveal A New Mechanism of Action with A Catalytic Dyad and Structural Twists. Sci Rep V. 11 3056 2021.
ISSN: ESSN 2045-2322
PubMed: 33542397
DOI: 10.1038/S41598-021-82660-Z
Page generated: Sun Jul 13 21:19:44 2025
ISSN: ESSN 2045-2322
PubMed: 33542397
DOI: 10.1038/S41598-021-82660-Z
Last articles
Cu in 7KF6Cu in 7KWO
Cu in 7KF8
Cu in 7KF7
Cu in 7KBT
Cu in 7K66
Cu in 7JRP
Cu in 7FB9
Cu in 7JRO
Cu in 7F0V